Cellular processes are dynamic and time-dependent
Many important dynamic cellular events and processes, such as cell differentiation, proliferation, migration, signaling, apoptosis, and morphological alterations of cells can be studied using live-cell imaging1. Cellular processes are often non-linear and their progression can occur over a very broad span of timescales, from minutes to days and weeks (Table 1).
Table 1 | Examples of applications of live-cell imaging and their duration.
| Application | Duration |
|
Cell culture monitoring |
Hours to several days |
|
Cell motion detection |
Minutes, hours, days |
|
Cellular structural alterations (e.g. morphological changes) |
Minutes, hours, days |
|
Cell growth/viability/proliferation |
Days |
|
Cell migration (e.g. chemotaxis assay) |
Minutes, hours, days |
|
Cell behavioral dynamics (e.g. wound healing assay) |
Usually up to 24 hours |
|
Dynamic complex assembly and interaction between cells |
Minutes, hours, days |
|
Cellular responses to environmental cues |
Minutes, hours, days |
Endpoint assays vs long-term live-cell imaging
Traditionally, endpoint assays have been used as a principal approach to characterize cell behavior. This provides a very static overview of a dynamic system, where the events occur in a non-linear fashion. Thus, sampling only at a singular time point could lead to misinterpretation of the results. For example, as illustrated in Figure 1, the dynamics of events are characterized by a bell-shaped curve and would not be accurately understood with only a one-time point of sampling.
As a result, the endpoint assay approach misses the opportunity to capture the dynamic cellular behaviors and responses, which could provide invaluable insights into biological processes under investigation, such as a deeper understanding of disease mechanisms and responses to treatments. Imaging live cells over time provides a more complete picture of dynamic biological processes through greater spatial and temporal resolution. Being able to learn about the kinetics of cellular reactions (e.g. when and how fast something happens) is instrumental to a better understanding of biological mechanisms.
There are several major benefits of being able to quantify transient cellular phenotypic responses, including:
- Optimization of time points for endpoint studies (assays);
- Clarifying results from previous endpoint studies that often arise from dynamic reversible processes that operate under precise temporal and spatial control;
- Characterization of adaptive responses;
- Determination of accurate scheduling and dosing regimens;
- Provision of additional data points of biological events collected over a time series;
- Facilitating more robust quantitative analysis from fewer specimens.
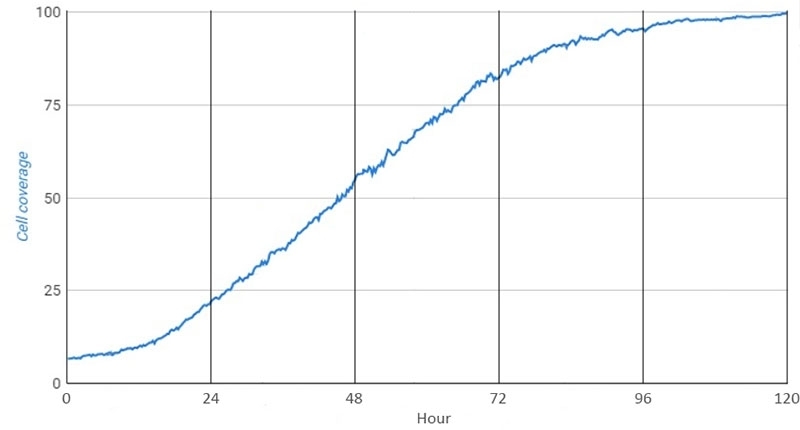
Researchers across different disciplines now routinely use time-lapse live-cell imaging to gain a more detailed understanding of cellular behavior. For instance, the graph in Figure 2 shows three different phases of cell confluence monitoring, where there is an initial lag phase (cell growth is about to start), followed by a log phase (cell growth is at its peak), and finally, a plateau phase (cell growth is halted due to e.g. reduced space or nutrient availability). In addition, we could use the information from the raw data to also determine the speed of cellular growth, which could be very insightful information, especially in situations when trying to optimize the medium for cell growth or trying to understand the influence of any other variables on cell growth (such as pharmacological or genetic manipulations). In many other applications of time-lapse imaging, the rate of changes in the parameters observed can provide critical knowledge about the biological processes, as depicted in the examples below.
Long-term live-cell imaging has become an invaluable analytical tool and a routine methodology in many fields of life and physical sciences, including biology, neurobiology, pharmacology, developmental biology, and many other related biomedical research disciplines3. It provides an unprecedented opportunity to accurately observe the dynamics of cellular behavior, often in response to external manipulations.
Applications of time-lapse live-cell imaging
There are many applications of time-lapse live-cell imaging in research and pre-clinical drug discovery, including the following:
- Colony Detection
- Cell confluence
- Cell culture monitoring
- Cell motion detection
- Cellular structural alterations (e.g. morphology analysis)
- Cell growth/viability/proliferation
- Cell migration (e.g. chemotaxis assays)
- Cell behavioral dynamics (e.g. would healing assays)
- Dynamic complex assembly and interaction between cells
- Cellular responses to environmental cues
- Cytotoxicity studies
- Phenotypic characterization of drug effects
- Pharmacokinetic studies (e.g. antiviral screening)
- Pharmacodynamic studies (e.g. drug toxicity assays)
Examples of time-lapse live-cell imaging
Cell confluence
Non-invasive cell confluence assay is required in many fields of biomedical research, including regenerative medicine, genetic engineering, and single-cell analysis. Monitoring cell confluence is essential for maintaining healthy cell cultures and preserving the native phenotypes. The two examples illustrated in Figures 1 and 2 show the applications of live-cell imagining in capturing cell confluence over a period of several days.
Collective cell migration: scratch assay
Cell motility is an essential component of many physiological (e.g. tissue regeneration) and pathophysiological processes (e.g. cancer or neurodegeneration). The movement is controlled by many internal and external signals, which together contribute to highly dynamic cell-cell and cell-substrate interactions. Being able to quantify the spatio-temporal dynamics of cell motility is crucial in order to understand cell behavior, which can only be achieved using time-lapse live-cell imaging. Scratch assays are a commonly used method of quantifying cell migration and identifying drugs affecting wound closure. Using time-lapse live-cell imaging methods, scratch assay parameters of wound closure can be easily monitored in real time and over prolonged periods of time. The initial step involves producing a “scratch” in a monolayer of cells and subsequently imaging living cells at regular time intervals for up to 24 hours to monitor cell migration, thereby being able to quantify cells’ rate of migration. This method is utilized for studies of cell-cell interactions, while fatefully reproducing events that occur during in vivo wound healing process4.
Drug discovery and development: pharmacokinetic and pharmacodynamic studies
Live-cell imaging plays an important role not only in the basic science that leads to the isolation and characterization of the proteins and environmental factors involved in the disease development, but also in drug discovery, optimization, and characterization at both the single-cell level, and subsequently, on tissues and organs inside animal models.
The discovery of new therapies for a particular disease often begins with identifying the drugs which can ameliorate the cellular malfunctions implicated in the disease5. Often, the diseases may cause particular phenotypic and behavioral changes at the cellular level, such as in various forms of cancer, including pancreatic, which can be studied using live-cell imaging6.
Quantitative in vitro imaging allows the evaluation of drug efficacy and drug response in complex biological systems. Real-time image-based analysis of drug response upon target activity and pathophysiology in vitro may accelerate drug development timelines, reduce costs, provide novel mechanistic insights into adaptive responses, and increase clinical predictivity when applied appropriately to relevant model systems.
An in vitro system that can test the dynamics of drug effects in a more systematic way can save time and costs in drug development. Long-term live-cell imaging assays can be used in the assay development stage of drug discovery to determine the endpoint assay to be used in a high-throughput screen to get an idea of how it would perform in vivo.
Drug actions on the human body depend strongly on pharmacokinetics, which describes drug absorption, distribution, and elimination over time. Single-cell imaging over time is playing an increasingly important role in the investigation of the pharmacokinetics of drugs or drug delivery systems in various diseases, especially cancers7.
Moreover, understanding the dynamic processes of cells provides an enhanced picture of drug pharmacology, enabling more reliable predictions of clinical responses earlier in the process. The insights about pharmacodynamics (such as cytotoxicity) gained from live-cell imaging over time enable better predictions about which drugs will be successful in clinical development to reduce late-stage attrition8.
Thus, tracking simple cellular parameters over time, such as phenotypic changes, or the number of live cells in culture wells following the treatment (such as chemotoxicity assays for cancer treatment) can be easily achieved using imaging of live cells over prolonged periods of time. In addition, live-cell imaging can provide information about the rates of changes in observed variables over time.
Conclusions
Related Products
There are currently no products tagged to this resource.